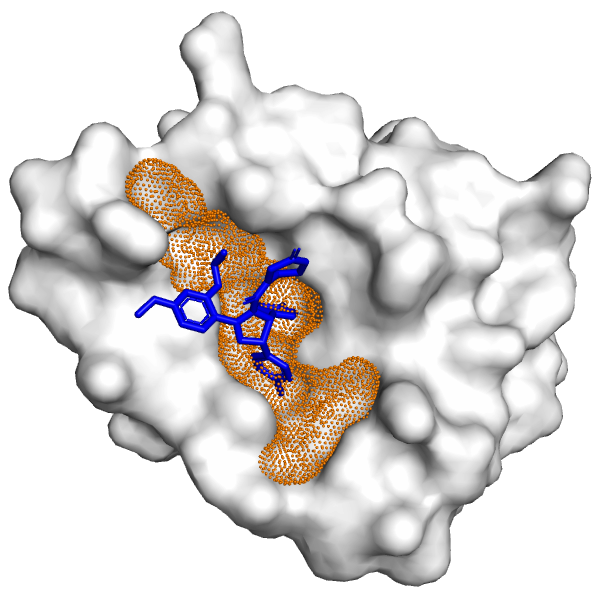
Prediction of ligand binding sites using improved blind docking method with a Machine Learning-Based scoring function - ScienceDirect
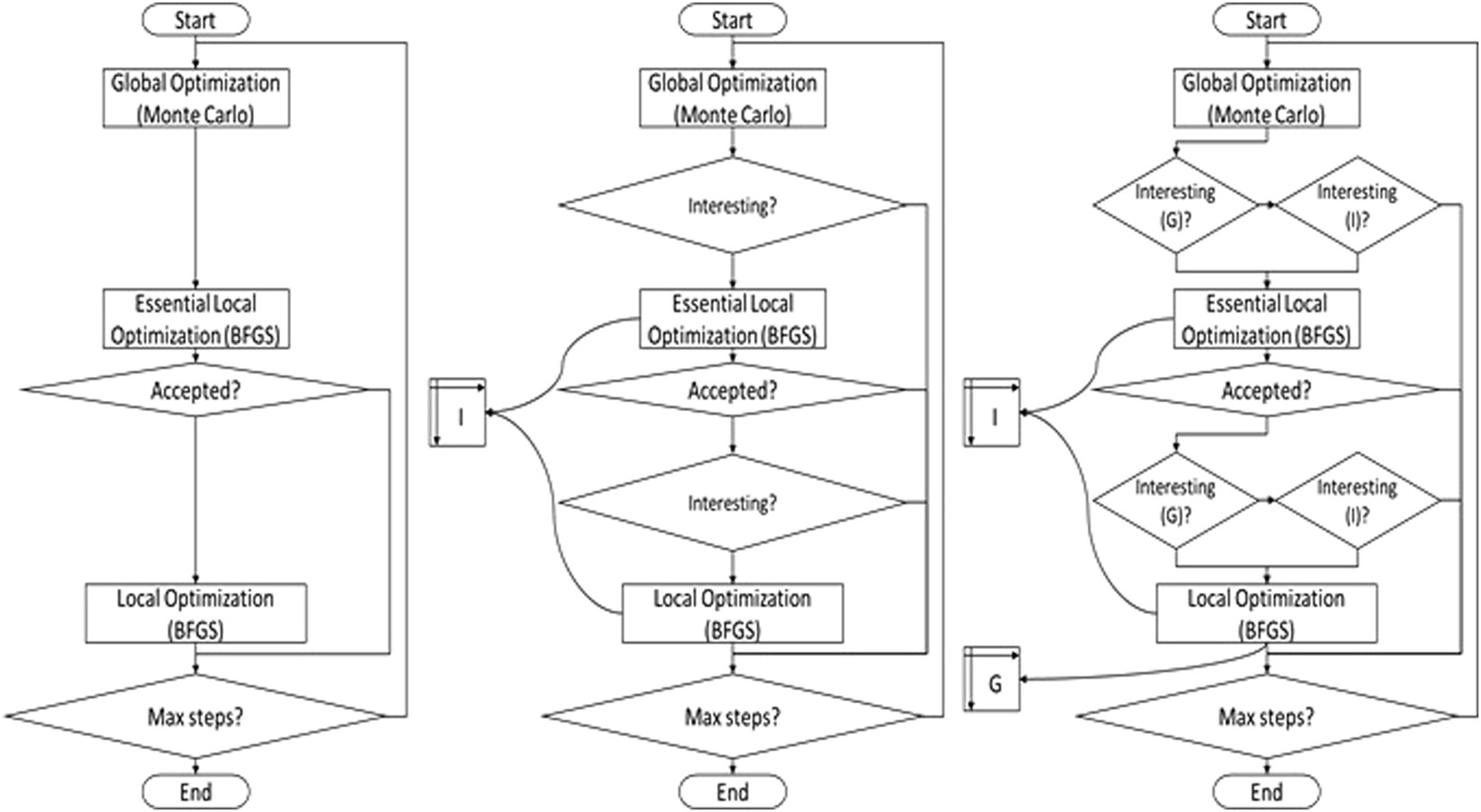
Protein-Ligand Blind Docking Using QuickVina-W With Inter-Process Spatio-Temporal Integration | Scientific Reports

Schematic representation of CoBDock blind docking workflow. The docking... | Download Scientific Diagram

Blind docking to the whole cavity of the hERG ion channel (A). Focused... | Download Scientific Diagram

Molecular docking in organic, inorganic, and hybrid systems: a tutorial review | Monatshefte für Chemie - Chemical Monthly

Fully Blind Docking at the Atomic Level for Protein-Peptide Complex Structure Prediction. | Semantic Scholar
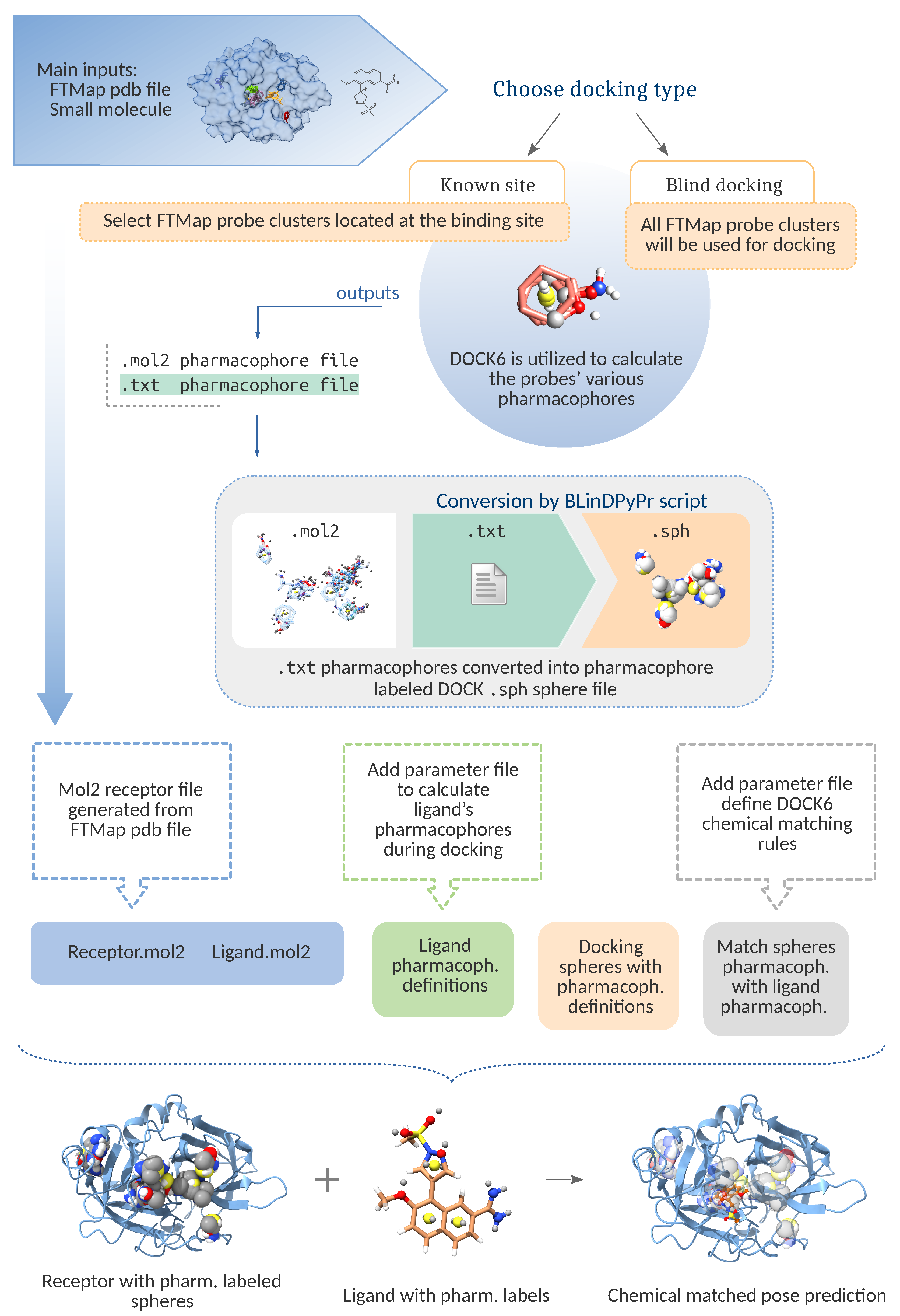
Molecules | Free Full-Text | Improving Blind Docking in DOCK6 through an Automated Preliminary Fragment Probing Strategy

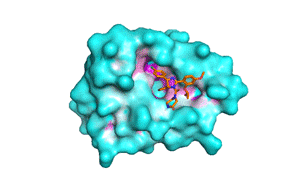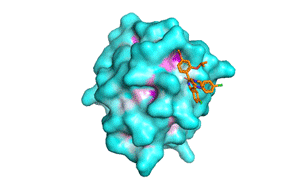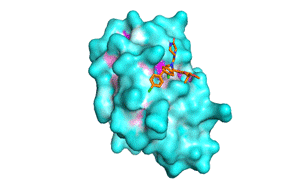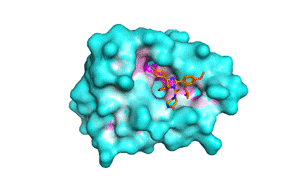
Blind docking at exhaustiveness 32 (a) Different binding conformations... | Download Scientific Diagram

Fully Blind Docking at the Atomic Level for Protein-Peptide Complex Structure Prediction - ScienceDirect
Improving accuracy and efficiency of blind protein-ligand docking by focusing on predicted binding sites

Grid box usage in docking: (A) blind docking with a grid box of size:... | Download Scientific Diagram
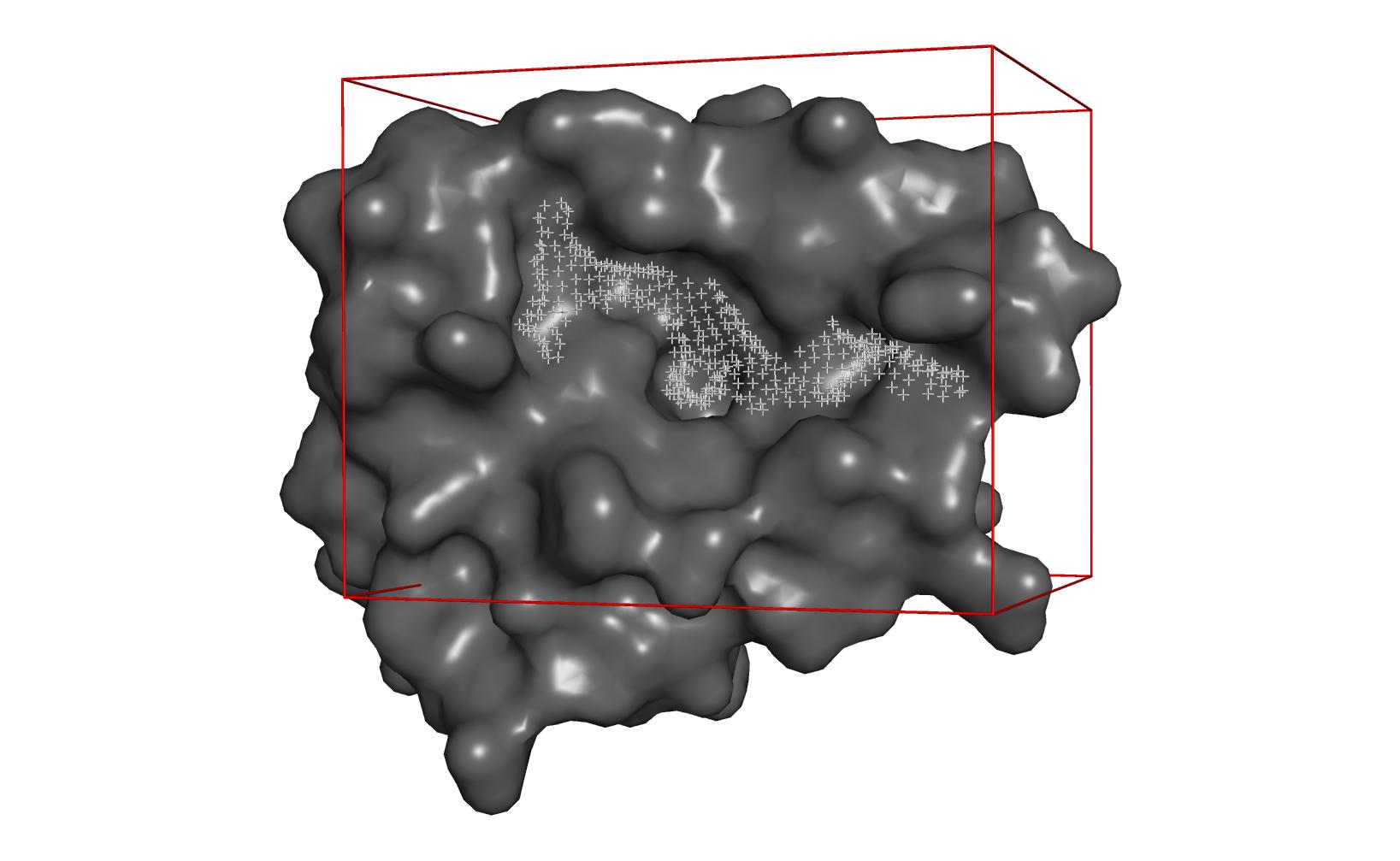


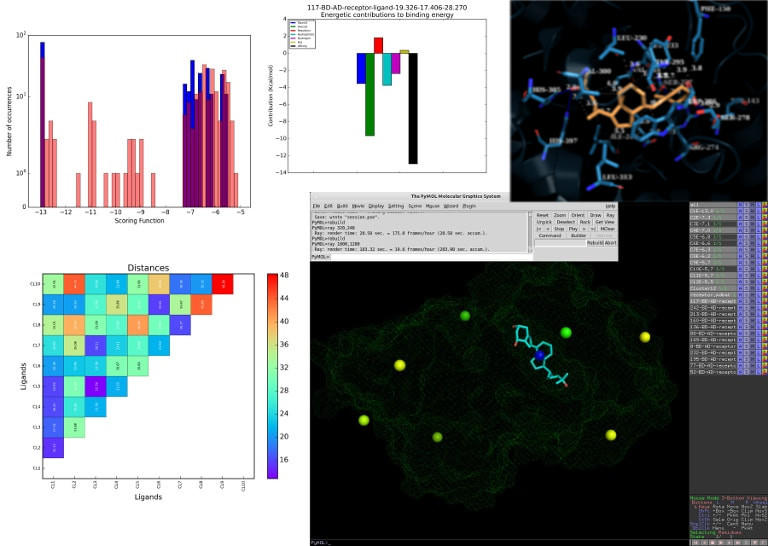
![PDF] DSDP: A Blind Docking Strategy Accelerated by GPUs | Semantic Scholar PDF] DSDP: A Blind Docking Strategy Accelerated by GPUs | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/b3f9ed5db9a0280d8c604ce18337bae8587c37d1/4-Figure1-1.png)